Isolation and identification of protease producing bacteria
Divyansh Rana1 , Mukesh Kumar Rana2 , Sheetal Shirodkar31Student, Amity Institute of Biotechnology, Amity University, Noida, Uttar Pradesh, India
2Principal Scientist, Division of Genomic Resources, ICAR-NBPGR, Pusa Campus, New Delhi, India

3AssistantProfessorII, Amity Institute of Biotechnology, Amity University, Noida, Uttar Pradesh, India
Abstract - Microbes are utilized in numerous industrial applications in a number of ways that exploit their own metabolic capabilities to aid human benefits. Proteases, the enzymatic product of bacteria, act as an integral component in a wide variety of industrial applications including pharmaceutical, food, leather and detergent formulation. Proteases are ubiquitous as they are foundinplants, animals, fungi, bacteria and viruses. Microbial sources of proteases, particularly bacteria, are of particular interest in industry. The isolation and identification of protease producing bacteria, hence, is the most basic pre-requisite in this regard for the success of any industry based on protease utilization. After their isolation from different sources, bacteria are routinely identified by morphological and biochemical characterization assays. Newer molecular techniques now allow bacterial species to be identified with authenticity by their genetic sequences. Different sources of bacteria for protease production have been exploited but soil samples, tannery and food processing industries including kitchen waste forms the basis of this report, wherein the procedures and protocols for their isolation and identification are discussed.
Key Words: Isolation,Protease,Identification,Biochemical characterization,Morphologicalassay
1.INTRODUCTION
Proteasesareagroupofenzymesthatcatalysetheprocessof proteolysisinwhichproteinsarebrokendownintosingle amino acids or smaller polypeptides. Proteases work to simplify proteins by disintegrating the peptide bonds by hydrolysispresentwithinthem.Proteaseenzymesalsohold apivotalroleinbiologicalfunctions,likethesimplificationof proteins into amino acids, their digestion followed by cell signaling(Kingetal.,2014;Shenetal.,2006).Proteasesalso causeachangebydisruptingthefunctionofaproteinand breakingitdowntoitssimplercomponents,whichleadsto the activation of a signal or a particular function in a biologicalpathway.Proteasesoccupyanimportantposition with respect to applications in commercial as well as physiological industries. Proteases act as an integral component in a wide range of industrial applications including pharmaceutical, food, leather and detergent formulation (Mohen et al., 2005). Also, of all the total enzymespresentglobally,proteasesaccountfortwothirds and about quarter of the total global enzyme produced
(Kumaretal.,2002).Proteasesarepresentinallorganisms including plants, animals, fungi, bacteria, and viruses. Differentsourcesofmicrobialproteasesfromwhichtheyare isolatedandidentifiedmostlyincludevariousbacteriaand fungi.Mostproteasesforcommercialutilization,areisolated from bacteria belonging to the genus Bacillus. Bacillus megaterium, Bacillus stearothermophilus, Bacillus thuringiensis,Bacilluspumilus,BacillussubtilisandBacillus stearothermophilus produce thermostable proteases. Besides Bacillus species, Clostridium histolyticum and variousStreptococcussp.,producecysteineproteasesand Vibriochlorae,PseudomonasaeruginosaandStreptomyces griseus produce metalloproteases (Gupta et al., 2002; Lakshmietal.,2008).Theproteasesobtainedfrommicrobial sourcesholdapivotalroleinindustrialapplicationssinceit leadstoagreatreductioninproductioncosts(Sawantand Nagendran, 2014). The production of these protease enzymesinmicrobesdependsontheculturemediatypeand compositionusedformicrobialgrowth,suchastheratioof carbontonitrogen,occurrenceofametabolizablesugarsuch as glucose etc. (Gupta et al., 2002). Besides, protease productiondependsuponthephysiologicalfactorssuchas theoptimumpH,optimumtemperature,inoculationmedia andtimeofincubation(Suppiahetal.,2012).Evenbiological characteristicslikethegenomeofanindividualcanleadto changeinthebiochemicalbehaviorofthemicrobeandaffect the production of metabolite (Varela et al., 1996). Commercially, optimization of culture media using numerousmediaconstituentsduringfermentationleadsto formationofacost-effectivemetabolicyield(Hameedetal., 1999). Also, till date no particular media is established to maximize the production of any enzyme because the genomicdiversitypresentinmicrobescauseseachmicrobial strain to exhibit a unique environment for maximizing enzymaticproduction.Hence,itisquiteessentialtoperform aseriesofexperimentsforinvestigatingthemetaboliteor enzymeproductionpatternunderdifferentenvironmental aspects from new isolated microbial strains to achieve maximumproduction(Prakashametal.,2006).Theprotease producing bacterial identification is the next step after successful isolation and is performed using various morphological and biochemical assays, supplemented by specifictestslikeantibioticinhibitionandserotypingtests. Recent research led to development of newer molecular diagnostic methodologies which allow microbial strain identification using their genetic sequences, directly from thesourcesample.Underbiochemicalcharacterization,the
International Research Journal of Engineering and Technology (IRJET) e-ISSN: 2395-0056

Volume: 10 Issue: 07 | Jul 2023 www.irjet.net
classification of protease producing bacteria can be done based on cellular structure, metabolism, or differences in cellular components. By performing graham’s staining techniquebacteriacanbecharacterizedbasedonstructure of cell walls possessed by them and on combining morphology with Gram-staining, the protease producing bacteriacanfurtherbeclassifiedasGram-positiveandGramnegativecocciandGram-positiveandGram-negativebacilli. Duetoadventofbiotechnologythatledtoadvancementsin molecularbiologyandcomputationalscience,theaccurate informationonphylogenyofaparticularbacterialstrainis possibleinnearfuture.
The literature is replete with information on isolation, purification, and characterization of bacterial strains for proteaseproductionfromdifferentsources(Banerjeeetal., 1999;JooandChang,2005;Sreejaetal.,2013),howeverthe proteaseproductionbybacterialisolatesfromsoil,kitchen wasteandtannerystillstandaslacunae.Theprimeobjective of the present report is to try to gather information on isolationandidentificationofproteaseproducingbacteria fromthesesources.
2. GENERAL TECHNIQUES
Throughouttheglobe,theprogressinbiotechnologyhasled toutilizingmicrobesforaidinghumanenvironment.Forthe successful usage of these microbes their isolation as well identificationisnecessary.Sincetimeimmemorialisolation and identification is being conducted. However due to industrializationadrasticchangeinmethodologyhasbeen observed that has led to isolation and identification of microbesinsimplewayswithlesstime.Inthereviewbelow thegeneralisolationandidentificationtechniquesusedby researchersallaroundtheworldarediscussed.
General techniques in isolation of bacteria
Advancements in biotechnology led to the discovery of culturingmicro-organismsinordertoidentifythemicrobes ofinteresttoaidhumanneeds.Toidentifythesemicrobes, from samples containing high microbial content, like soil, wastes, industrial effluents, leather etc. isolation is the preliminarystep.Forisolatingamicrobephysically,ithasto be cultured as a pure culture. Pure culture is defined as a culturepreparedinlaboratorythatconsistsofonlyasingle speciesofaparticularmicrobe.Ifanyoneexpectstoisolate foraparticularmicrobeofinterest,themicrobialcultureas well as the isolation techniques need to be modified and geared towards that microbe. For example, if a particular microbeisintoleranttoair,itcanbeisolatedusinganaerobic isolationmechanisms only. Therefore,thedevelopmentof pure culture entirely depends on the knowledge of the optimal oxygen requirements, nutritional needs and temperatureforgrowth.
There are two universally accepted methodologies for isolatingmicrobes:
1. Streaking by isolation of microbes on agar plates
1. Pour plate method of isolation
Streaking by isolation of microbesonagar plates involves continuous serial dilutions of sample containing microbes untilthecellsexistinaverylowdensity,sothatthesingle cells obtained thereafter can be isolated spatially to form individualbacterialcolonies.Onthecontraryinpourplate methodforisolation,youseriallydilutethemicrobial rich samplesufficiently,andthenaddittoamoltencooledagar inadish.Theisolatedbacterialcellswillleadtoformationof individualbacterialcoloniesontheagar.Theonlydrawback tothismethodologyforisolationisthatifthemoltenagar hot,itwilldenatureandkillthemicrobesandifitistoocold, it will form a big lump in the petri dish. So due to these reasonsstreakingmethodispreferredasityieldsbacterial coloniesofinterestontheagarsurface.
General techniques in identification of bacteria
After successful isolation of the bacterial colony, its identificationisalsonecessaryinordertoidentifytowhich strainthemicrobebelongsto.Todothisfollowingmicrobial identificationtechniquesareused:
Macroscopic and microscopic features
Macroscopicidentificationusesphysicalappearancewhich includes shape, size, color, and smell. For describing microscopicfeaturesforidentificationuseofamicroscopeis required to infer whether they are rod, cocci, or spiralshaped,haveflagellaorbudsformedandhaveafilamentous hyphaornot.
Staining for identification
Stains in cytology enable easier and clear visualization of microbesunderamicroscope.Stainingisperformedusing differentstainingtechniques.
Gram Staining - Gramstainingisanimportanttestdonefor bacterialidentification.Thispurplestaintestisbasedonthe usageofacrystalvioletdye(Colco,2005) Itdistinguishes microbesbasedonthepropertiesoftheircellwalls.Grampositivemicrobespossessthickpeptidoglycanlayer,sothey retainthecrystalvioletstainwhilegram-negativemicrobes possessathinpeptidoglycanlayer,sotheydonotretainthe stain.
Endospore Staining - Itisanidentificationtechniquethat involves application of a stain to the microbial sample for checking the presence of spores using a primary stain malachitegreen.Duetothefactthatnotallbacteriaproduce spores, this is a useful technique for identification. The various types of endospores identified are free, central, swollenandsubterminalendospores.(Husseyetal.,2012)
International Research Journal of Engineering and Technology (IRJET) e-ISSN: 2395-0056

Volume: 10 Issue: 07 | Jul 2023 www.irjet.net
Fungi staining - Fungal staining can be helpful for identifyingthefungalelementsandcharacteristicsoffungal speciemicrobesbasedoncellwallstaining.
Lactophenol cotton blue –thisstainnormallycolorsthe carbohydrateportionspresentincellwallsofthefungus blue (Heritageetal.,1996)
Periodic-acid Schiff stain (PAS stain) –thisstaincolors thecarbohydratesalongwithsomeothercomponentsof fungalcellwallproducingamagentacolorinlivingfungi only.
Grocot’s methenamine silver stain –thisstaincolorsthe cellwalloffungibrowntoblack,butdoesnotdistinguish whetherthefungiislivingordead.(Nassaretal.,2006)
Biochemical testing for identification
Catalase testing - Unidentified bacterial species can be identified using the catalase activity test in which oxygen bubbles appearwhen hydrogen peroxide is added to a bacterial species cultured on a slide. (Mahon et al., 2011; McLeodandGordonetal.,1923)
Oxidase testing - Using oxidase test, the identification of bacterial isolates is achieved by determining the cytochrome-coxidase(CCO)activity.Ifcytochrome-coxidase exists, it oxidizes a reagent called tetramethyl-pphenylenediamine forming a purple-colored product. If cytochrome-c oxidase is absent, the reagent remains in reducedstateformingacolorlessproduct.(McFaddin,2000)
Substrate utilization tests - Substrate utilization tests is usually performed to identify bacteria using a variety of substrates made from carbon and nitrogen sources. The utilizationofsubstratesbybacterialspeciesleadstocolor changes in substrates on incubation with bacteriawhich thengeneratesakeyofhowthesubstrateisutilized.Thekey isthencomparedwithpre-existingsubstrateusepatternsof microbespresentonvariousdatabasestogeneratealistof bacterialspecies.(Campbelletal.,2003)
Chemical/analytical identification
Fatty acid profiling and chemo-profiling - Inbacteria,fatty acidsareanintegralcomponentoftheircellmembranes,and dependingonthespeciesofbacteriadifferentcombinations offattyacidsareproduced.Profilingofthesefattyacidsis done, and it is used toidentify the bacterial strain by comparing them to known profiles. Besides this, bacteria alsoproducevarioussecondarymetaboliteslikeantibiotics, antioxidants and immunosuppressive compounds. On the basis of type of secondary metabolites produced, differentbacterial strains are identified using chemo profilingmethodologyofidentification.(Kumarietal.,2008)
Modern methodologies
p-ISSN: 2395-0072
Thedrawbacksoftheoldtechniquesarethattheyidentify thebacteriathatarecultured in vitro. Additionally,someof the bacterial strains exhibit distinct biochemical characteristicsthatcouldnotbeidentifiedusingprimitive methodologies as they did not fit in any genus or species. The advent of biotechnology has led to introduction of modernmethodsforidentifyingbacteriawhichrevealeven theminutedifferencesbetweenorganisms.
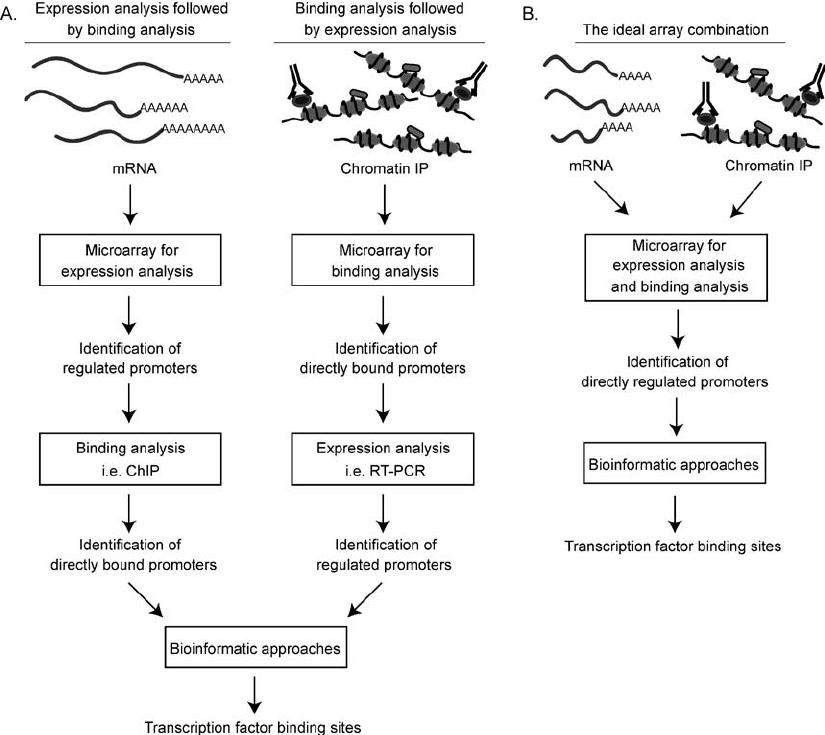
Microarray-Based Identification - Microarray-based identification of microbesis based on hybridization of amplified DNA sequences of microbes to specific oligonucleotide probes that vary from species to species. Further,eachoftheseoligonucleotideprobesaremadeupof differentdyesthatshowfluorescenceonhybridization.Once hybridized,thefluorescentregionsonthesehybridscanbe comparedtostandardhybridsandspeciescanbeidentified (Caoetal.,2011).(Figure-1)
Figure-1: Microarray based identification methodology using specific oligonucleotide probes varying species to species
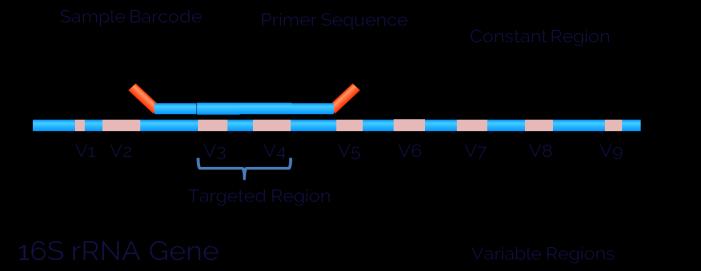
Next generation sequencing - In case of all organisms possessing DNA including various bacterial strains, the presence of 16S rRNA gene (Figure below) acts as a conserved component of the transcription machinery. (Petrosinoetal.,2009).Thisgeneactsasatargetgenefor sequencingDNAfrombacterialstrainshavingthousandsof differentspecies.Fortargetingtheconservedregionsin16S rRNAgene,universalprimersforPCRhavebeendesignedto amplifythetargetgenefromasinglesample.Besidesthis, the 16S rRNA gene is composed of both variable and conservedregions.Whiletheroleoftheconservedregionis to do universal amplification, the sequencing of variable regionsof16SrRNAgeneallowsdifferentiationofdifferent bacterialeadingtotheiridentification(Figure-2).
International Research Journal of Engineering and Technology (IRJET) e-ISSN: 2395-0056

Volume: 10 Issue: 07 | Jul 2023 www.irjet.net p-ISSN: 2395-0072
isolationandidentificationofproteaseproducingbacteria fromdifferentsourceshasbeenthoroughlyscrutinizedand findingsofvariousstudiesareprovidedhereunderdifferent headingsdependinguponthetypeofsourceselected.
1. Isolation and identification of protease producing bacteria from soil samples
Different NGS methods such as 454 pyrosequencing, Illumina,SOLiD,IonTorrent,andsingle-moleculereal-time (SMRT) circular consensus sequencing equipment from Pacific Biosciences (Lu et al., 2009) and Oxford Nanopore have provided more pace and deep analytic power in identificationofmicrobes(Nichollsetal.,2019).
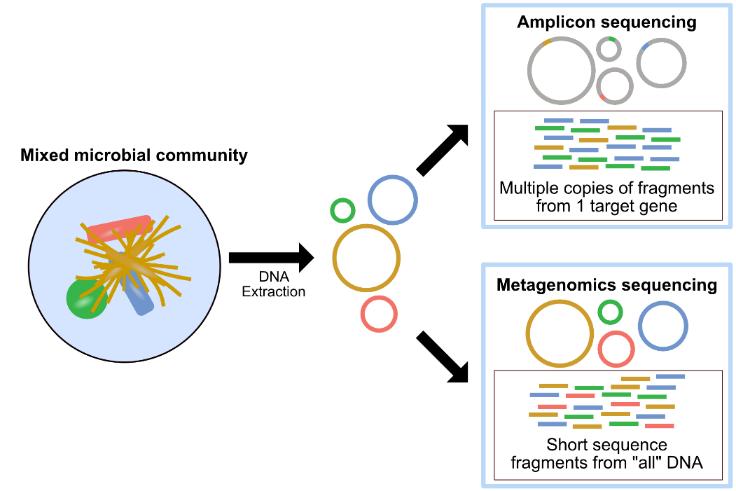
Metagenomics - Metagenomicsinvolves the genomic analysisofmicrobesthatcannotbecultured(unculturable) by direct extraction of their DNA, and consequently comparingitwithwell-establishedribosomalsequencesof microbes, so that the unseen microbial population can be identifiedandpurified.Metagenomicsisalsointerpretedas the study of entire microbial genomes present in mixed communities,whichallowsthemtobeidentifiedatstrainlevel resolution. Due to the fact that 16S or 18S or ITS sequencingisnotspecificenoughtoperformcomprehensive microbiomeexperiments,theuseofmetagenomicsenables theidentificationoftherarestofmembersintheseunseen microbialcommunities.(Kumarietal.,2013)(Figure-3)
(SujathaandSubash,2017)investigatedtheextractionand characterization of an extracellular protease from thebacterialstrainof Bacillus subtilis isolated fromtermite infestedsoilsample.Amongthefiveproteaseproducersthat wereisolated,oneoftheisolateswasselectedforresearch. Thisisolationwasrelatedto Bacillus subtilis onthebasisof identification using various biochemical characterization techniquesand16SrRNAgenesequencing.
1.1 Isolation of bacteria
Termiteinfestedsoilsamplewascollected,andisolationwas performed using serial dilutions followed by spread plate technique.Thesamplewasfirstdiluteduptoten-foldsand 0.1mlfrom10-4 dilutionwasintroducedonskimmilkagar plateandincubatedatroomtemperaturefor48hours.Post incubation,thebacterialstrainsproducingthelargestclear hydrolytic zone were selected and then sub-cultured on nutrientagarplates.Anisolate‘M1’whichshowedmaximum caseinolyticactivity,wasselectedforpurificationtoobtain protease.
1.2 Identification of bacteria
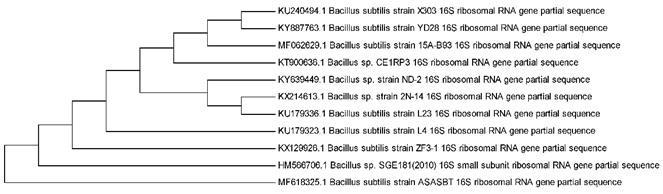
DNA extraction from the collected sample was performed usingtheCTABmethod(Ausubeletal.,1992).Furtherthe 16srRNAsequencingwasperformed.Postsequencingthe sequencewascomparedwithapre-existinggenesequence on various databases using BLAST program on NCBI webpage (Castresana, 2000), evolutionary history was traced (Saitou and Nei, 1987) and phylogenetic tree was madeusingMUSCLE3.7(Edgar,2004)asshowninFigure1. Basedonthe16SrRNAgenesequencingmethodologyfor identification,phylogeneticanalysiswasperformed,andit indicatedthattheisolatedM1bacteriawas Bacillus subtilis. (Figure-4)
3. METHODOLOGY
Lotofworkhasbeencarriedoutintheisolation,purification and characterization of bacterial strains for various purposes. However, literature is scanty as far as protease producingbacteriaarespecificallyconcerned.Literaturefor
International Research Journal of Engineering and Technology (IRJET)

Volume: 10 Issue: 07 | Jul 2023
1.3 Characterization of protease enzyme from bacterial strain
Qualitative and quantitative assays of protease from isolated Bacillus subtilis
The bacteria Bacillus subtilis was inoculated in a culture mediumandtheculturewassubjectedtocentrifugationto obtainenzymesource. For quantitative assay, theactivity of enzyme protease in the culture of the bacteria was determinedbythemethodologyofGaurav et al.(Pantetal., 2015)For qualitative determination of protease, the proteaseenzymethatwasextractedwasculturedonacasein agarplatetoconfirmthepresenceofprotease.Onincubation at room temperature the clear zone was measured and presence of the enzyme was confirmed (Pant et al., 2015, OlajuyigbeandAjele,2005) Bacillus subtilis bacterialstrain wasfurthersubjectedtodifferenttypesofcultureconditions inordertoidentifytheoptimumconditionsformaximizing the production of protease. Protease production was estimated at a wide pH range of 5.0 to 10.0, at varying temperaturesfrom20°Cto 70ºCandcarbonandnitrogen sourceslike dextrose, K2HPO4, sucrose, lactose, starch, gelatin,maltose,Na2PO4,ureaandbeefextracts(Abbasand Leila, 2011). Ammonium sulphate fractionation method based on salting out principle was used to purify the obtainedproteasesshowninTable1andTable2showsthe purificationprofileusingDEAEcellulosemethod(Simpson, 2004,Roe,2004,Sookkheoetal.,2000).
Table-1: Purification of protease produced from Bacillus subtilis using ammonium sulphate precipitation methodology
Table-2: Purification of DEAE-Cellulose purified ammonium sulphate precipitated protease from the microbe
2. Isolation and identification of protease producing bacteria from tannery samples
(Pertiwiningrum et al., 2017) isolated and identified proteaseproducingbacteria fromsolidandliquidtannery waste.Theisolationofproteaseproducingbacterialstrains was performed using a sample of solid and liquid wastes taken from different waste reservoirs (three liquid waste reservoirs and one solid waste reservoir) from leather manufacturing units followed by their identification using biochemicalcharacterizationassays.
2.1 Isolation of bacteria
Numerous wastewatersampleswerecollectedfromareas adjoining tanneries. Solid and liquid waste samples were obtained from four varying reservoirs. The first reservoir had tannery waste that was not processed, the second reservoir had tannery waste which was processed by aeration, and the third reservoir had tannery waste post processing. The solid waste samples were taken from the solidwastereservoir(sludge).SampleT1,T2andT3were isolatedfromliquid wasteoffirst,secondandthirdwaste storage respectively, while sample T4 was isolated from fourth solid waste storage. For isolation of bacteria, 1mL sample was taken in test tubes. and by serial dilution the concentrationwasreducedfrom10-1 to10-10.Then,100µm of10-8,10-9 and10-10dilutionsweretaken,andintroduced onskimmediausingpourplatemethod,followedbytheir incubationat27°Cfor3days.Thetestwaspositivedueto formationofaclearzonearoundthebacterialcoloniesand one colony was selected. Post purification the isolated samplesweregrownonagarslants,followedbyincubation at 27°C for 72 hours, and stored at 5°C (Rahayu, 1991; Miyamotoetal.,2002).Theisolatethatshowedthehighest proteolyticenzymeactivitywasisolateT2.
2.2 Identification of bacteria
Various morphological and biochemical characterization assays including gram staining, motility test, catalase test, macroscopicandmicroscopicmorphology,elevationconvex, spherical shape, and white-coloured colonies were performedfortheidentification.Furthercharacterizationof isolateT2basedonmicroscopicmorphologyinterpretedthey were rod shaped, pink, gram-negative, non-motile and catalasepositive.Basedontheseresults,thebacterialisolate T2waspredictedtobelongtogenus Bacillus.
2.2.1 Measuring clear zone of hydrolysis diameter
AgarmediumswithdifferentpHtreatmentrangingfrom7to 11 were added with one drop of pre-culture followed by incubationfor3days.Postgrowthofbacterialcolonies,their clear-zonediameterwasmeasuredwithcalipers(Schmidtet al., 2011). On performing the data analysis of clear zone diametersofbacterialisolateT2inthemediawithdifferent pHdeterminedthattheclearzonediameterofpH7wasnot verydifferentfromthediametersofpH8,pH9andpH12 (Table3).Onthecontraryitwasdifferentfromtheoneswith pH10and11(Table3).Thebestclearzonediameterwas produced in medium with pH 11 which meant bacterial isolatesT2grewbestinmediumcontainingpH 11.Hence bacterial isolate T2 belonged to the alkaliphilic group therebymakingtheenzymeproducedbybacteriaisolatesT2 asanalkalineproteaseenzyme.
Table-3: Measuring the colony diameter, clear-zone diameter and proteolytic index of bacterial isolate T2
measuringthe proteolytic indexofan enzyme,it becomes easytodeterminethebacterialabilityforproteaseenzyme activityinagoodquality(Syafieetal.,2013).Asshownin Table 1, the proteolytic index of bacterial isolate T2 remainedunaffectedevenontreatmentwithdifferentmedia ordifferentpHappliedtothemedia.
2.3.2 Enzyme activity test
SincebacterialisolateT2wasanalkalineproteaseenzyme itsactivitytestincludestheblankandsamplemeasurement as well as the standard measurement of tyrosine. To each testtubefirst0.5mlbuffersolutionhavingvaryingpHfrom7 to12wasaddedfollowedbyadditionof0.5mLofcaseinas substrate and 1 mL of distilled water for blank measurement,1mLofthesampleforsamplemeasurement and1mLoftyrosineforstandardmeasurement.All these testtubeswerethenincubatedatroomtemperaturefor10 minutes, followed by addition of 1ml 10% trichloroacetic acidandincubationatroomtemperature,andfilteredwith Whatmanfilterpaper.0.75mlofthefiltratewastakenina testtube,and2.5mL0.5Mofsodiumcarbonatewasadded along with 0.5 mL folin’s reagent and kept at room temperaturefor15minutes.Theabsorbancewasexamined and noted at 578 nm (Nadeem et al., 2007; Ahmed et al., 2008).
2.3.3 Protease Enzyme Characterization
ThevaryingpHaddedtotheculturemediumhadnoeffect onthetotal activityorthespecific activityoftheprotease enzyme(Table4).SincebacterialisolatesT2couldgrewbest in medium containing pH 11, highest enzyme activity and specific activity could be seen at pH 11. Therefore, it was inferredthattheoptimumenzymeactivityofisolateT2was atpHof11.Onthecontrary theenzymeactivitybasedon varyingtemperature(40–60oC)hadnodifferenceassuch (Table 5). However, the optimum enzyme activity for T2 bacterialisolatewasat40°C.
Table-4: Bacterial isolate T2 enzyme activity and specific activity of enzyme in different pH
**Allthemeanshavingdifferentsuperscriptsinthesame indicatesignificantlydifferent(P<0.05)andns:nonsignificant(P>0.05)
2.3 Characterization of protease enzyme from bacterial strain
2.3.1 Proteolytic index measurement

Proteolyticindexofanenzymecanbedeterminedusingthe diameter of the clear zone and the diameter of bacterial colonyasitistheratioofclearzonearea’sdiametertothe diameter of bacterial colonies (Baehaki et al., 2011). By
**ns:non-significant(P>0.05)
Table-5: Bacterial isolate T2 enzyme activity and specific activity of enzyme on treatment with a wide range of temperatures
Table-6: The physiological and biochemical characterization of bacterial isolate B1 and B2 form kitchen waste samples
Type of test
isolate B1
isolate B2 Gramstaining Grampositive Grampositive
Endosporetype Centralplaced Centralplaced Morphology
colony Rodshaped bacteria Rodshaped bacteria
**ns:non-significant(P>0.05)
3. Isolation and identification of protease producing bacteria from kitchen waste and food processing industries

(Gill et al., 2016) collected samples from kitchen waste which consisted of vegetables and fruit peels, fruit fibers, thrown away cooked food items and various food grains. PDB(proteindegradingbacteria)strainswereisolatedusing nutrientagarmedia byenrichment techniquefollowed by theiridentificationusingBergey’smanualofdeterminative bacteriology.
3.1 Isolation of bacteria
Theisolationof bacterial strainswasperformedusingthe enrichment technique. Primarily in this technique the samplesareincubatedin125mlflasksfilledwith50mlof nutrient agar medium (g/L). The composition of this nutrientagarmediawas5gmofpeptone,3gmofyeastand beefextractinequalproportions,15gmofagar,5gmofNaCl andpHmaintainedat7.4.Thetypeofenrichmenttechnique used for isolation of bacterial strains was serial dilution. Usingserialdilution,1gramofkitchenwastesampleperml was added to 9 ml distilled water and dilution was performed up to factor of 10-6 under a laminar airflow to maintainasepticenvironment.Then0.1mlofsamplefrom each dilution was plated on nutrient agar followed by incubationatroomtemperaturefor2days.Bacterialisolates growinginnutrientagarslantsweremaintainedat40oC.
3.2 Identification of bacteria
Both the bacterial isolates (B1 and B2) obtained by screening were identified through morphological and biochemical characterization of cultures according to Bergey’s Manual (Bergey et al., 1974) (Table-6). These isolatesalsounderwentmicroscopicobservationlikecolony size, pigmentation, form, elevation and colony color etc. (Table-7).Onthebasisoftheseidentificationmethodologies isolateB1wasfoundtobe Bacillus megaterium andB2was foundtobe Bacillus subtilis
**✔:positivereactions, ✘:negativereactions,d:dubious reactions
S.No. Bacterial Isolates (B1 and B2)
Morphology observed
1) B1bacterial isolates Slightlyirregularshaped havinganundulatedmargin
2) B2bacterial isolates Dry,flatandirregular shapedhavinglobatetypeof margins
3.3 Characterization of protease enzyme from bacterial strain
3.3.1 Screening of protease producing microbes

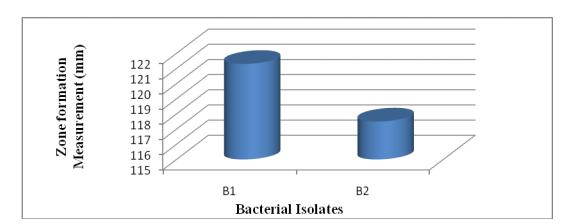
Usingmodifiedbasalmedium(MM),thequalitativeenzyme activityassaywasdetermined.Thecompositionofmodified basalmedium(g/L)was14gmofagaralongwith6.2g/L skim milk protein, 5 g/L of casein as substrate, 1 gm of glucoseand2.5gmofyeastextract.Whenthebasalmedium wasintroducedwithbacterialisolates,formationofaclear zoneofhydrolysisinthemediumaroundthewell,suggested thatproteaseactivityoccurred(Perezetal.,2009).Twoout ofsixisolatesshowedformationoftransparentcircularzone aroundbacterialcoloniesonskimmilkagarplateandgelatin agarplatewhichsuggestedthepresenceofprotease(Sinha et al., 2013; Alnahdi, 2012). Similar methodology for screeningusingskimmilkandgelatinagarwasusedearlier by (Abirami et al., 2011; Geethanjali and Subhash, 2011; SevincandDemirkan,2011;Smitaetal.,2012;Sinhaetal., 2013).Basedontheappearanceofclearzonesofhydrolysis two bacterial isolates having protease production capabilitieswereselected.Thesehalozonesoftwobacteria aregivenbelowinTable-8andFigure-5.
Figure-5: Zone formation measurement of the bacterial isolates from kitchen waste
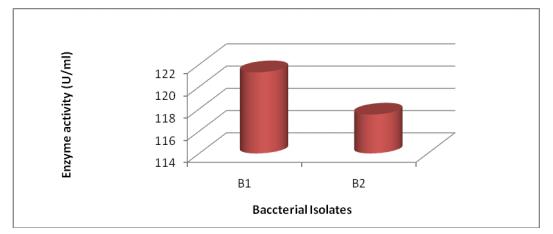
3.3.2 Quantitative assay for protease activity
Using the modified methodology proposed by (Folin and Ciocalteu, 1927)protease enzyme assay was performed which exhibited as in Figure-6, that maximum enzyme activityofbacterialisolateB1wasmorethanB2.Sosuitable changes were made in the basal medium to find the best sourceformaximumenzymeproduction.
4. RESULT AND DISCUSSION
Theresultsofthestudyconductedby(SujathaandSubash, 2017)showedthatthemaximumenzymeactivityrecorded at48hoursofincubationatroomtemperaturewas154.13 µ/ml. The optimum pH of 7 and temperature at 40 °C showedmaximumproductionofenzymeprotease.Thebest carbon and nitrogen sources for production of proteases werestarchandgelatin,respectively.Theisolatedprotease wassubjectedtopurificationinfourdifferentstepsnamely ammonium sulphate precipitation, followed by dialysis, DEAE-CelluloseandfinallysephadexG-100chromatography whichhada4.01-foldincreaseinthespecificactivityofthe enzyme protease (86.51 µ/mg) and 23.73 % increase in recoverypercentageoftheenzyme.
Parallelly, using the same methodology as performed by (SujathaandSubash,2017)researchwasalsoconductedby (Vishnuetal.,2016)and(Vijitraetal.,2019)onisolationand identificationofproteaseproducingbacteria.(Vishnuetal., 2016) isolated protease producing bacterial strains from marine soil in Tamil Nadu. Using biochemical assays for identification,itwasidentifiedas Pseudomonas fluorescens
Pseudomonas fluorescens wasinoculatedingelatinagarfor protease enzyme production. The enzyme extract was purifiedusingammoniumsulphatefractionationand85% saturation was obtained. The enzyme activity was determined as 7.5 µ/ml after incubation for 1 day. The maximum extracellular protease production was found at roomtemperature(37oC)andthespecificactivityofenzyme was 49.02 µ/mg. Similarly, studies by (Vijitra et al., 2019)identifiednineteenbacterialisolateshavingprotease producing capabilities using 16S rRNA gene sequencing technique.Seventeenisolatesoutofnineteenwereregarded as Bacillus spp. Based on biochemical assays only two isolates were identified as Staphylococcus sp. and Enterobactersp.Bacillusthuringiensis wasfoundtoshowthe highest enzyme activity of 3.72 ± 0.08 µ/mg at optimum conditionsoftemperature65°CandpH8post30minutes incubationin0.05MPBSbufferwith1%caseinassubstrate. Theresultsofthestudyconductedby(Pertiwiningrumetal., 2017) indicated that the morphology of the T2 isolate showedcircular,convexelevation,pink,gram-negative,nonmotile, gelatin negative and catalase positive. The highest recordedenzymewasachievedatapHof11withenzyme specific activity as 43,19±1,69 µ /mg and total activity as 45,18±1,77 µ /ml and at optimum temperature of 40°C enzymeactivitywas54,02±1,89µ/mlandspecificactivity was 51,65±1,8 µ /mg. The protease enzyme activity from precipitatedammoniumsulphateshowedahigherresultof (75,8 µ /ml) as compared to rough protease. Further the study conducted by (Gill et al., 2016) resulted in the selectionof,twoeffectiveproteindegradingbacteriaoutof the six bacterial isolated strains. Both these isolates underwentthequalitativeassayandquantitativeassay,and the protease enzyme activity was measured. The results from protease enzyme assay indicated that, isolate B1 exhibited the maximum enzyme activity of 121.3 µ/mL at 37oCafterthreedaysincubation.Onthecontrary,isolateB2 exhibitedlowestenzymeactivityof117.5µ/ml.Therefore, glucose was replaced by different sugars in the basal medium and fructose was found as the best source for productionofmaximumprotease125µ/mL)by Bacillus sp. (Sevinc and Demirkan, 2011). Similar methodology for isolationandidentificationasdiscussedbywasadoptedby (SonyandPotty,2016)forisolatingandidentifyingprotease producing bacteria from food processing industries. According to their study protease producing bacteria was isolated from soil and wastewater samples of a bakery Kollamregion,India.Proteaseproductionwasexaminedby seriallydilutingthesamples,followedbyintroducingthem ongelatinagarplatesat37oCfor2days.Total87dissimilar bacterial colonies showed protease activity which was confirmedbytheoccurrenceofzonearoundtheisolates.Out ofthese87bacterialcolonies,27showedoccurrenceofclear zone of hydrolysis around them. Further, out of these 27 isolates,eightwereselectedbasedondiameterofclearzone. These eight isolates were further identified using biochemicalcharacterizationsandbiochemicalidentification using Biomerieux VITEK 2 system. These isolates were
identifiedas Proteusmirabilis,Cedeceadavisae,Enterobacter asburiae and Staphylococcus intermedius. Proteaseenzyme activitywasthenmeasuredusingstandardmethodologyand enzymewaspurifiedandusedasatoolforwasterecyclingin foodprocessingindustries.
5. FUTURE PERSPECTIVE
Proteaseenzymes find application in diverse end-use markets,suchasdetergentsformulations,pharmaceuticals anddiagnostics,foodprocessingandotherindustriessuchas textile,animalfeedandchemical.Proteasesareanintegral component of all the industrially significant enzymes involvedinmostofthecellularandphysiologicalprocesses. Sincetheyareinvolvedinphysiologicalprocesses,theyare abundantlyfoundinmicrobes,animalsandplants.However, proteases from microbes are of particular commercial interest and are preferred because of their fast growth, reduced space requirement for their growth and easy amenability and credibility to genetic manipulation. Since time immemorial protease enzymes are being extensively utilized in the food, dairy and detergent formulation industries.Apartfromthis, theapplicationofproteases in therapeuticshasbeengrowingrapidlyoverthelastdecade. There is a rekindled interest in using proteases as target enzymes for the development of drugs to treat the uncontrollablefataldiseaseslikecancer,AIDSandmalarial parasites.Also,therecentdiscoveriesandmethodologiesuse genetically manipulated small molecule activated (SMA) protease enzymes which leads to activation of apoptotic enzymecaspase(Grayetal.,2010)whichactsasanewway forcontrollingtheproteaseactivityinhumansandcanbe used in diagnostics. Further the exploitation of these proteolytic activities of proteases are opening up the vast potential for the future in diseased tissues as they offer a newwayforthedevelopmentofsite-specificdrugs(Ersteret al.,2012)aswellasimagingtumors(Jiangetal.,2004)
The discovery of new sequencing techniques has allowed deeper insights in studying the evolutionary relationship amongproteases.Microbialproteaseswhich,hitherto,were not amenable to biotechnological interventions are now becoming so. This understanding of evolutionary relationshipsamongproteaseswouldleadtoisolationand identification of novel strains of bacteria producing these proteases.Therefore,theexploitationofbiologicalresources in order to generate and extract microbes producing proteasessuitedfordifferentindustrialapplicationsaiding humans is considered as the most promising future alternatives.Geneticengineeringandproteinengineeringis nowbecomingpossibletoalterthepropertiesofproteases andwouldlead toimprovedstrains of bacteria producing these proteases that were never made in nature and that wouldmeettherequirementsofthemultitudeofprotease applications.

Withtheadvancementsinmicrobiologyandbiotechnology, wecanvisualizethattheutilizationofproteasesinthenear futurewillbeamulti-disciplinarytaskduetowhichhumans couldattainmanydramaticsuccesses.
6. CONCLUSION
This report is mainly focused on microbial proteases, however special emphasis is given on the isolation, purificationandidentificationoftheseproteaseproducing bacteria.Proteasesfrommicrobesplayaverycrucialrolein numerous industrial applications such as detergent formulation,diagnosticsandtherapeutics,agriculturaland chemicalindustriesandarebecominganalternativeinthe nearfuture.Theutilizationofmicrobesinordertoproduce enzymes like proteases has far too many advantages and recentlyhasbecomethepreferredmodeforproductionof enzymes. However, the costs related to the production of theseenzymesisquitehighandcostsofitsprocurementare evenmore,hencethepresentreporthighlightsthepotential to obtain protease producing bacteria at minimal costs which could be alternative for industrial and commercial use.
(SujathaandSubash,2017)investigatedtheextractionand characterization of an extracellular protease from termite infestedsoilsample. Basedon16SrRNAgenesequencing they identified the isolate to be Bacillus subtilis Protease producingbacteriahavebeenisolatedandidentifiedfrom tannery waste ( Pertiwiningrum et al., 2017). Based on variousmorphologicalandbiochemicalassays,theprotease producing bacteria from tannery waste was predicted to belongtogenus Bacillus whosemorphologyforidentification showed it to be circular, convex elevation, pink, gramnegative,non-motile,gelatinnegativeandcatalasepositive. Foodprocessingindustries &kitchenwastehasalsobeen usedtoisolateproteaseproducingbacteria(Gilletal.,2016; Sony and Potty, 2016) 49 observed that two isolates had proteaseproducingcapabilitieswhichfurtherunderwentfor qualitativetestingforidentificationandquantitativetesting for determining the protease enzyme activity. Apart from this,extensiveresearchisbeingconductedgloballyhunting forastableenzymehavinghighefficiencyandeffectiveness in various classes of applications. According to the recent advancementsinscienceandtechnologyarapidlygrowing trend has been observed in commercial applications of proteasesandtheuseofmicrobialsourcestoisolatenovel proteaseenzymesforaidinghumanneeds.Theprogressand developmentinbiotechnologyisnowofferingaconstructive position for the production of protease enzymes which wouldcontinuetoacceleratetheirutilizationstoachievea sustainableenvironment.

REFERENCES
[1] King,JohnV.;Liang,WenguangG.;Scherpelz,KathrynP.; Schilling,AlexanderB.;Meredith,StephenC.;Tang,W.-J.
Molecularbasisofsubstraterecognitionanddegradation byhumanpresequenceprotease.Structure 22,996–1007 (2014).
[2] Shen, Yuequan; Joachimiak, Andrzej; Rosner, Marsha Rich;Tang,W.-J.Structuresofhumaninsulin-degrading enzymerevealanewsubstraterecognitionmechanism. Nature 443,870–874(2006).
[3] Mohen FN, Dileep D, D. D. Potential application of protease isolated from Pseudomonas auriginosa. BiotechnolInd 8,197–203(2005).
[4] KumarA,SachdevA,BalasubramanyamSD,SaxenaAK,L. A.Optimizationofconditionsforproductionofneutral and alkaline protease from species of Bacillus and Pseudomonas.IndJMicrobiol 42,233–236(2002).
[5] Gupta R, Beg QK, L. P. Bacterial alkaline proteases: molecularapproachesandindustrialapplications.Appl MicrobiolBiotechnol 59,15–32(2002).
[6] Lakshmi P, Rao PS, Devi P, L. N. Production of a halotolerant alkaline protease by exiguobacterium acetylicumMTCC9115.TheEcoscan 3,135–138(2008).
[7] Sawant, R.; Nagendran, S. Protease: An enzyme with multipleindustrial applications.World J.Pharm.Sci. 3, 568–579(2014).
[8] SuppiahS,SendeshkannanK,PrabakaranP,RajkumarG, Y. & N. Purification and characterization of alkaline protease from Lysini Bacillus fusiformis. J Biochem Technol 4,561–564(2012).
[9] Varela H, Ferrari MD, Belobradjic L, Weyraueh R, L. M. Effectofmediumcompositionontheproductionofanew Bacillus isolate of protease with promising unharing activity. World J Microbiol Biotechnol 12, 643–645 (1996).
[10] HameedA,KeshavarzT,E.C.Effectofdissolvedoxygen andpHontheproductionofextracellularproteasesfrom a new isolate of Bacillus subtilis K2. J Chem Technol Biotechnol 74,5–8(1999).
[11] PrakashamRS,SubaRaoCH,S.P.Greengramhuskand inexpensivesubstrateforalkalineproteaseproduction by Bacillus sp in solid state fermentation. Bioresour Technol 97,1449–1454(2006).
[12] Sreeja SJ, Malar JPW, Joseph SFR, Tiburcius S, I. G. Optimization of cellulase production by Bacillus altitudinisAPSMSUandBacilluslicheniformisAPS2MSU, gut isolates of fish Etroplus suratensis. Int J Adv. Res Technol 2,123–132(2013).
[13] Banerjee CW, Sani RK, Azmi W, S. R. Thermostable alkaline protease from Bacillus brevis and its
characterization as a laundry detergents additive. ProcessBiochem 35,213–219(1999).
[14] Joo HS, C. C. Production of protease from a new alkalophilic Bacillus sp. I-312 grown on soybean meal: optimizationandsomeproperties.ProcessBiochem 40, 1263–12701(2005).
[15] R,C.‘GramStaining’.Curr.Protoc.Microbiol. 0,(2005).
[16] Hussey,Marise;Zayaitz,A.EndosporeStainProtocol.Am. Soc.Microbiol. 4,123–154(2012).
[17] Killington,J.H.E.G.V.E.R.A.IntroductoryMicrobiology. CambridgeUniv.Press100(1996).
[18] Hauser,C.MayoClinicGastroenterologyandHepatology BoardReview.CRCPress108(2005).
[19] NassarA,ZapataM,LittleJV,S.M.Utilityofreflexgomori methenaminesilverstainingforPneumocystisjirovecii onbronchoalveolarlavagecytologicspecimens:Areview. Diagn.Cytopathol 34,719–723(2006).
[20] Mahon, C. R., D. C. Lehman, and G. M. Textbook of diagnosticmicrobiology.(W.BSaundersCo.,2011).
[21] McLeod, J. W., and J. G. Catalase production and sensitiveness to hydrogen peroxide amongst bacteria: with a scheme for classification based on these properties.J.Pathol.Bacteriol. 26,326–331(1923).
[22] BiochemicalTestsforIdentificationofMedicalBacteria. (LippincottWilliamsandWilkins,2000).
[23] CampbellCD,ChapmanSJ,CameronCM,DavidsonMS,P. J. A rapid microtiter plate method to measure carbon dioxideevolvedfromcarbonsubstrateamendmentssoas todeterminethephysiologicalprofilesofsoilmicrobial communitiesbyusingwholesoil.ApplEnv.Microbiol. 69, 3593–3599(2003).
[24] KumariP,BijoAJ,MantriVA,ReddyCR,J.B.Fattyacid profilingoftropicalmarinemacroalgae:ananalysisfrom chemotaxonomicandnutritionalperspectives.(2008).
[25] CaoB,LiR,XiongS, etal.UseofaDNAmicroarrayfor detection and identification of bacterial pathogens associatedwithfisheryproducts.ApplEnv.Microbiol. 77, 8219–8225(2011).
[26] Petrosino J F, Highlander S, L. R. A. Metagenomic pyrosequencingandmicrobialidentification.Clin.Chem. 55,856–866(2009).
[27] LuT,StrootPG,O.D.Reversetranscriptionof16SrRNA tomonitorribosome-synthesizingbacterialpopulations intheenvironment.Appl.Environ.Microbiol. 75,4589–4598(2009).
[28] NichollsSM,QuickJC,TangS,L.N.Ultra-deep,long-read nanopore sequencing of mock microbial community standards.Gigascience 8,43(2019).
[29] KumariP,BijoAJ,MantriVA,ReddyCR,J.B.Fattyacid profilingoftropicalmarinemacroalgae:ananalysisfrom chemotaxonomic and nutritional perspectives. Phytochemistry 86,44–56(2013).
[30] Sujatha A and Subash A. Isolation and screening of extracellularproteaseproducedbyBacillussubtilisfrom termite soil. in Life Science: Research, Practices and Application for Sustainable Development. 708–713 (2017).
[31] AusubelF,BrentR,KingstonR,MooreD,S.J.and&J,S. Currentprotocolsinmolecularbiology.NewYork-Greene Publ.Assoc.Wiley-Interscience(1992).
[32] J, C. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol.Evol. 17,540–552(2000).
[33] M, S. N. and N. The neighbor-joining method: a new methodforreconstructingphylogenetictrees.Mol.Biol. Evol. 4,406–425(1987).
[34] 34. RC, E. Muscle: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5,113(2004)
[35] PantG,PrakashA,P.J.andB.S.Production,optimization andpartialpurificationofproteasefromBacillussubtilis. J.TaibahUnivSci 9,50–55(2015).
[36] JO, O. F. and A. Production dynamics of extracellular proteasefromBacillussp.Afr.J.Biotechnol. 4,776–779 (2005).
[37] Abbas AS, L. J. Effect of culture conditions on the production of an extracellular protease by Bacillus sp. Enz.Res.1–7(2011).
[38] RJ, S. Purifying proteins for proteomics: A Laboratory ManualColdSpringHarborLaboratoryPress,NewYork. (2004).

[39] S, R. Protein purification techniques: A practical approach.OxfordUniv.Press.SecondEd.NewYork 38, 130–131(2004).
[40] SookkheoB,SinchaikulS,P.S.andC.S.Purificationand characterizationofthehighthermostableproteasesfrom Bacillusstearothermophilus.Prot.Exp.Purification. 20, 142–151(2000).
[41] A., P., D., A. F., A., F. N. & Rochijan, A. Isolation and identification of bacterial protease enzyme of leather waste.J.Indones.Trop.Anim.Agric. 42,33–41(2017).
[42] K., R. Isolation and activity testing enzyme. PAU food Nutr. Univ. Gadjah Mada Press. Yogyakarta. 4, 23–42 (1991).
[43] Miyamoto,K.,H.Tsujibo,E.Nukui,H.Itoh,Y.&Inamori, K. and Y. Isolation and characterization of the genes encodingtwometalloproteases(MprIandMprII)froma marine bacterium, Alteromonas sp. strain. Biosci. Biotecnol.Biochem. 66,416–421(2002).
[44] Schmidt, J., M. Müsken, T. Becker, Z., Magnowska, D. Bertinetti,S.Möller,B.,Zimmermann,F.W.Herberg,L.J. and & Häussler, S. The Pseudomonas aeruginosa chemotaxismethyltrasferaseCheR1impactsonbacterial surfacesampling.PLoSOne 6,(2011).
[45] Baehaki,A.,R.danA.B.Isolationandcharacterizationof proteases from Indralaya soil swamp bacteria, South Sumatera.J.Teknol.FoodInd. 22,10–16(2011).
[46] 46 Syafie,Y.,S.T.danA.P.TheutilizationofAspergillus sp.andRhizopussp.proteaseatdifferentconcentration inunhairingphaseoftanningonphysicalqualityofsheep leatherandliquidwaste.FarmBull. 37,198–206(2013).
[47] Nadeem, M. J.I. Qazi, S. B. and Q. S. Studies on commerciallyimportantalkalineproteasefromBacillus lichniformis N-2 isolated from decaying organic soil. Turk.J.Biochem. 32,171–177(2007).
[48] Ahmed,S.A.,R.A.Al-domany,N.M.A.E.-S.&Saleh.,H.H. R.andS.A.Optimization,immobilizationofextracellular alkalineproteaseandcharacterizationofits enzymatic properties.Res.J.Agr.Biol.Sci. 4,434–446(2008).
[49] Gill, S. S., Shrivastav, A. & Jana, A. M. Isolation and Identification of Protease producing Bacteria through BiodegradationofProteincontentofKitchenWastesin Gwalior, India. Int. J. Curr. Microbiol. Appl. Sci. 5, 204–211(2016).
[50] Bergey D.H., B. R. E. and G. & N.E. Bergey’s Manual of DeterminativeBacteriology.8thedition.WilliamsWilkins Co.,Balt.USA15–36(1974).
[51] PerezBorla,O.,Davidovich, L.A., andR.&S.I.Isolation andcharacterizationofproteolyticmicroorganismsfrom freshandfermentedcabbage.LWTFoodSci.Technol. 43, 298–301(2009).
[52] Sinha,P.,Singh,R.K.,Srivastva,R.,S.&R.,andTiwari,P. Characterization and optimization of alkaline protease enzymeproducedbysoilbornebacteria.TrendsLifeSci. 2,2319–4731(2013).
[53] Alnahdi, H. S. Isolation and screening of extracellular proteasesproducedbynewisolatedBacillussp.J.Appl. Pharma.Sci. 2,71–74(2012).
[54] Abirami, V., Meenakshi, S.A., K., K., Bharathidasan, R., Mahalingam, R. & and Panneerselvam, A. Partial Purification and Characterization of an Extracellular ProteasefromPenicilliumjanthinellumandNeurospora crassa.Eur.J.Exp.Biol. 1,114–123(2011).
[55] A., G. S. and S. Optimization of protease production by Bacillussubtilisisolatedfrommidgutoffreshwaterfish Labeorohita.WorldJ.FishMar.Sci. 3,88–95(2011).
[56] E.,S.N.andD.ProductionofProteasebyBacillussp.N-40 IsolatedfromSoil andItsEnzymaticProperties. J.Biol. Environ.Sci. 5,95–103(2011).
[57] SmitaG.S.,R.P.andM.S.QuantificationandOptimisation ofBacterialIsolatesforProductionofAlkalineProtease. AsianJ.Exp.Biol.Sci. 3,180–186(2012).
[58] V., F. O. and C. On tyrosine and tryptophane determinations in proteins. J. Biol. Chem. 73, 627
650 (1927).
[59] VishnuPriyaVG,PreethiS,K.S.andR.B.N.G.Isolation and identification of protease producing bacteria from soil.Int.Res.J.Eng.Technol. 3,1362–1365(2016).
[60] Vijitra Luang-In, Manatchanok Yotchaisarn, Worachot Saengha,PiyachatUdomwong,S.D.andK.M.Proteaseproducing bacteria from soil in Nasinuan community forest, Mahasarakham province, Thailand. Biomed. Pharmacol.J. 12,587–595(2019).
[61] 61. Sony I.S., P. V. P. Isolation and identification of protease producing bacteria from food processing industries. Int.J.Curr.Microbiol.App.Sci. 5, 181–189 (2016).

[62] Gray, D., Mahrus, S. and Wells, J. Activation of specific apoptoticcaspaseswithanengineeredsmall-moleculeactivatedprotease.Cell 142,637–646(2010).
[63] Erster,O.,Thomas,J.,Hamzah,J.,Jabaiah,A.,Getz,J.,S.& T., Hall, S., Ruoslahti, E. and Daugherty, P. Site-specific targeting of antibody activity in vivo mediated by disease_associated proteases. J. Control. release 161, 804–812(2012).
[64] Jiang,T.,Olson,E.,Nguyen,Q.,Roy,M.,Jennings,P.and& Tsien, R. Tumor imaging by means of proteolytic activationofcell-penetratingpeptides.Proc.Natl.Acad. of.Sci. 101,17867–17872(2004).
