Listofcontributors
AnshikaAgarwal InsilicoResearchLaboratory,EminentBiosciences,Indore, MadhyaPradesh,India
SarahAlbogami DepartmentofBiotechnology,CollegeofScience,Taif University,Taif,SaudiArabia
NandadulalBairagi DepartmentofMathematics,CentreforMathematical BiologyandEcology,JadavpurUniversity,Kolkata,WestBengal,India
KrishnanBalasubramanian SchoolofMolecularSciences,ArizonaState University,Tempe,AZ,UnitedStates
SubhashC.Basak DepartmentofChemistryandBiochemistry,Universityof Minnesota,Duluth,MN,UnitedStates
EmilioBenfenati LaboratoryofEnvironmentalChemistryandToxicology,Istituto diRicercheFarmacologicheMarioNegriIRCCS,Milano,Italy
ApurbaK.Bhattacharjee DepartmentofMicrobiologyandImmunology, BiomedicalGraduateResearchOrganization,SchoolofMedicine,Georgetown University,Washington,DC,UnitedStates
AnushkaBhrdwaj InsilicoResearchLaboratory,EminentBiosciences,Indore, MadhyaPradesh,India;DepartmentofBioinformatics,ComputerAidedDrug DesigningandMolecularModelingLab,AlagappaUniversity,Karaikudi,Tamil Nadu,India
SumanK.Chakravarti MultiCASEInc.,Beachwood,OH,UnitedStates
PratimKumarChattaraj DepartmentofChemistry,IndianInstituteof TechnologyKharagpur,Kharagpur,WestBengal,India
SamratChatterjee ComplexAnalysisGroup,TranslationalHealthScienceand TechnologyInstitute,NCRBiotechScienceCluster,Faridabad,Haryana,India
xvi Listofcontributors
RamanaV.Davuluri DepartmentofPreventiveMedicine,DivisionofHealthand BiomedicalInformatics,NorthwesternUniversityFeinbergSchoolofMedicine, Chicago,IL,UnitedStates
TathagataDey CentreforInterdisciplinaryResearchandEducation,Kolkata, WestBengal,India;DepartmentofComputerScience&Engineering,Indian InstituteofTechnologyBombay,Mumbai,Maharashtra,India
AbhikGhosh IndianStatisticalInstitute,Kolkata,WestBengal,India
IndiraGhosh SchoolofComputational&IntegrativeSciences,JawaharlalNehru University,NewDelhi,Delhi,India
GiuseppinaGini PolitecnicodiMilano,DEIB,PiazzaLeonardodaVinci,Milano, Italy
LimaHazarika InsilicoResearchLaboratory,EminentBiosciences,Indore, MadhyaPradesh,India
GuangHu DepartmentofBioinformatics,CenterforSystemsBiology,Schoolof BiologyandBasicMedicalSciences,SoochowUniversity,Suzhou,P.R.China
ChiakangHung PolitecnicodiMilano,DEIB,PiazzaLeonardodaVinci,Milano, Italy
TajamulHussain BiochemistryDepartment,CollegeofScience,KingSaud University,Riyadh,SaudiArabia;CenterofExcellenceinBiotechnologyResearch, CollegeofScience,KingSaudUniversity,Riyadh,SaudiArabia
YanrongJi DepartmentofPreventiveMedicine,DivisionofHealthand BiomedicalInformatics,NorthwesternUniversityFeinbergSchoolofMedicine, Chicago,IL,UnitedStates
IshaJoshi InsilicoResearchLaboratory,EminentBiosciences,Indore,Madhya Pradesh,India
TaushifKhan ImmunologyandSystemsBiologyDepartment,OPC-Sidra Medicine,Ar-Rayyan,Doha,Qatar
RavinaKhandelwal InsilicoResearchLaboratory,EminentBiosciences,Indore, MadhyaPradesh,India
PawanKumar NationalInstituteofImmunology,ArunaAsafAliMarg,New Delhi,Delhi,India
ShivamKumar ComplexAnalysisGroup,TranslationalHealthScienceand TechnologyInstitute,NCRBiotechScienceCluster,Faridabad,Haryana,India
MinLi DepartmentofBioinformatics,CenterforSystemsBiology,Schoolof BiologyandBasicMedicalSciences,SoochowUniversity,Suzhou,P.R.China
JieLiao DepartmentofPathology,NorthwesternUniversityFeinbergSchoolof Medicine,Chicago,IL,UnitedStates
ClaudiuN.Lungu DepartmentofChemistry,FacultyofChemistryand ChemicalEngineering,Babes-BolyaiUniversity,Cluj,Romania;Department ofSurgery,FacultyofMedicineandPharmacy,UniversityofGalati,Galati, Romania
SubhabrataMajumdar AIVulnerabilityDatabase,Seattle,WA,USA;Bias Buccaneers,Seattle,WA,USA
RamaK.Mishra DepartmentofBiochemistryandMolecularGenetics,Feinberg SchoolofMedicine,NorthwesternUniversity,Chicago,IL,UnitedStates
ManjuMohan InsilicoResearchLaboratory,EminentBiosciences,Indore, MadhyaPradesh,India
AsheshNandy CentreforInterdisciplinaryResearchandEducation,Kolkata,West Bengal,India
AnurajNayarisseri InsilicoResearchLaboratory,EminentBiosciences,Indore, MadhyaPradesh,India;DepartmentofBioinformatics,ComputerAidedDrug DesigningandMolecularModelingLab,AlagappaUniversity,Karaikudi,Tamil Nadu,India;BiochemistryDepartment,CollegeofScience,KingSaudUniversity, Riyadh,SaudiArabia;BioinformaticsResearchLaboratory,LeGeneBiosciences PvtLtd,Indore,MadhyaPradesh,India
ShahulH.Nilar GlobalBloodTherapeutics,SanFrancisco,CA,UnitedStates
RanitaPal AdvancedTechnologyDevelopmentCentre,IndianInstituteof TechnologyKharagpur,Kharagpur,WestBengal,India
AditiPande InsilicoResearchLaboratory,EminentBiosciences,Indore,Madhya Pradesh,India
GuillermoRestrepo MaxPlanckInstituteforMathematicsintheSciences, Leipzig,Germany;InterdisciplinaryCenterforBioinformatics,LeipzigUniversity, Leipzig,Germany
DipankaTanuSarmah ComplexAnalysisGroup,TranslationalHealthScience andTechnologyInstitute,NCRBiotechScienceCluster,Faridabad,Haryana,India
DwaipayanSen CentreforInterdisciplinaryResearchandEducation,Kolkata, WestBengal,India
SanjeevKumarSingh DepartmentofBioinformatics,ComputerAidedDrug DesigningandMolecularModelingLab,AlagappaUniversity,Karaikudi,Tamil Nadu,India
ChillamcherlaDhanalakshmiSrija InsilicoResearchLaboratory,Eminent Biosciences,Indore,MadhyaPradesh,India
RevathyAryaSuresh InsilicoResearchLaboratory,EminentBiosciences,Indore, MadhyaPradesh,India
MuyunTang DepartmentofBioinformatics,CenterforSystemsBiology,School ofBiologyandBasicMedicalSciences,SoochowUniversity,Suzhou,P.R.China
GarimaThakur InsilicoResearchLaboratory,EminentBiosciences,Indore, MadhyaPradesh,India
XinTong DepartmentofPathology,NorthwesternUniversityFeinbergSchoolof Medicine,Chicago,IL,UnitedStates
MarjanVracko TheoryDepartment,Kemijskiins ˇ titut/NationalInstituteof ChemistryLjubljana,Slovenia
MarjanVrac ˇ ko NationalInstituteofChemistry,Hajdrihova19,Ljubljana, Slovenia;TheoryDepartment,Kemijskiins ˇ titut/NationalInstituteofChemistry, Ljubljana,Slovenia
ZeWang DepartmentofPharmaceuticalSciences,ZunyiMedicalUniversityat ZhuhaiCampus,Zhuhai,P.R.China
DanDanXu DepartmentofPathology,NorthwesternUniversityFeinbergSchool ofMedicine,Chicago,IL,UnitedStates
Guang-YuYang DepartmentofPathology,NorthwesternUniversityFeinberg SchoolofMedicine,Chicago,IL,UnitedStates
Elsevier
Radarweg29,POBox211,1000AEAmsterdam,Netherlands TheBoulevard,LangfordLane,Kidlington,OxfordOX51GB,UnitedKingdom 50HampshireStreet,5thFloor,Cambridge,MA02139,UnitedStates
Copyright©2023ElsevierInc.Allrightsreserved.
Nopartofthispublicationmaybereproducedortransmittedinanyformorbyanymeans, electronicormechanical,includingphotocopying,recording,oranyinformationstorageand retrievalsystem,withoutpermissioninwritingfromthepublisher.Detailsonhowtoseek permission,furtherinformationaboutthePublisher’spermissionspoliciesandour arrangementswithorganizationssuchastheCopyrightClearanceCenterandtheCopyright LicensingAgency,canbefoundatourwebsite: www.elsevier.com/permissions .
Thisbookandtheindividualcontributionscontainedinitareprotectedundercopyrightby thePublisher(otherthanasmaybenotedherein).
Notices
Knowledgeandbestpracticeinthisfieldareconstantlychanging.Asnewresearchand experiencebroadenourunderstanding,changesinresearchmethods,professionalpractices, ormedicaltreatmentmaybecomenecessary.
Practitionersandresearchersmustalwaysrelyontheirownexperienceandknowledgein evaluatingandusinganyinformation,methods,compounds,orexperimentsdescribed herein.Inusingsuchinformationormethodstheyshouldbemindfuloftheirownsafety andthesafetyofothers,includingpartiesforwhomtheyhaveaprofessionalresponsibility.
Tothefullestextentofthelaw,neitherthePublishernortheauthors,contributors,or editors,assumeanyliabilityforanyinjuryand/ordamagetopersonsorpropertyasamatter ofproductsliability,negligenceorotherwise,orfromanyuseoroperationofanymethods, products,instructions,orideascontainedinthematerialherein.
ISBN:978-0-323-85713-0
ForInformationonallElsevierpublications visitourwebsiteat https://www.elsevier.com/books-and-journals
Publisher: SusanDennis
AcquisitionsEditor: CharlotteRowley
EditorialProjectManager: KyleGravel
ProductionProjectManager: SujathaThirugnanaSambandam
CoverDesigner: GregHarris
TypesetbyMPSLimited,Chennai,India
Section1Generalsection
1Chemoinformaticsandbioinformaticsbydiscretemathematics andnumbers:anadventurefromsmalldatatotherealmof emergingbigdata3 SubhashC.Basak
1.1Introduction3
1.2Chemobioinformatics—aconfluenceofdisciplines?5
1.2.1Physicalproperty:colligativeversusconstitutive6
1.2.2Earlybiochemicalobservationsontherelationship betweenchemicalstructureandbioactivityofmolecules6
1.2.3Linearfreeenergyrelationship:themultiparameter Hanschapproachtoquantitativestructure activity relationship7
1.2.4Chemicalgraphtheoryandquantumchemistryasthe sourceofchemodescriptors9
1.3Bioifnormatics:quantitativeinforamticsintheageofbigbiology19
1.4Majorpillarsofmodelbuilding21
1.5Discussion24
1.6Conclusion27 Acknowledgment29 References29
2Robustnessconcernsinhigh-dimensionaldataanalysesand potentialsolutions37 AbhikGhosh
2.1Introduction37
2.2Sparseestimationinhigh-dimensionalregressionmodels39
2.2.1Startingoftheera:theleastabsoluteshrinkageand selectionoperator39
2.2.2Likelihood-basedextensionsoftheLASSO40
2.2.3Searchforabetterpenaltyfunction41
2.3Robustnessconcernsforthepenalizedlikelihoodmethods43
2.4PenalizedM-estimationforrobusthigh-dimensionalanalyses44
2.5Robustminimumdivergencemethodsforhigh-dimensional regressions46
2.5.1Theminimumpenalizeddensitypowerdivergence estimator47
2.5.2AsymptoticpropertiesoftheMDPDEunder high-dimensionalGLMs49
2.6Areal-lifeapplication:identifyingimportantdescriptorsof aminesforexplainingtheirmutagenicactivity51
2.7Concludingremarks54
Appendix:AlistofusefulR-packagesforhigh-dimensional dataanalysis55
Acknowledgments56 References56
3Fairness,explainability,privacy,androbustnessfor trustworthyalgorithmicdecision-making61
SubhabrataMajumdar
3.1Introduction61
3.2Fairnessinmachinelearning61
3.2.1Fairnessmetricsanddefinitions62
3.2.2Biasmitigationinmachinelearningmodels63
3.2.3Implementation66
3.3Explainableartificialintelligence67
3.3.1Formalobjectivesofexplainableartificialintelligence67
3.3.2Taxonomyofmethods69
3.3.3Doexplanationsservetheirpurpose?71
3.4Notionsofalgorithmicprivacy73
3.4.1Preliminariesofdifferentialprivacy74
3.4.2Privacy-preservingmethodology76
3.4.3Generalizations,variants,andapplications79
3.5Robustness81
3.5.1Adversarialattacks82
3.5.2Defensemechanisms83
3.5.3Implementations84
3.6Discussion84 References84
Section2Chemistry&chemoinformaticssection
4Howtointegratethe“smallandbig”dataintoacomplex adverseoutcomepathway?99
MarjanVra ˇ cko
4.1Introduction99
4.2Stateandreview101
4.3Bindingaffinitytoandrogennuclearreceptorevaluatedwith respecttocarcinogenicpotencydata104
4.4Conclusionandfuturedirections106 References111
5Bigdataanddeeplearning:extractingandrevising chemicalknowledgefromdata115 GiuseppinaGini,ChiakangHungandEmilioBenfenati
5.1Introduction115
5.2Basicmethodsinneuralnetworksanddeeplearning117
5.2.1Neuralnetworks117
5.2.2Neuralnetworklearning119
5.2.3Deeplearningandmultilayerneuralnetworks120
5.2.4Attentionmechanism123
5.3Neuralnetworksforquantitativestructure activityrelationship: input,output,andparameters124
5.3.1Input125
5.3.2Chemicalgraphsandtheirrepresentation125
5.3.3Output127
5.3.4Performanceparameters127
5.4Deeplearningmodelsformutagenicityprediction128
5.4.1Structure activityrelationshipandquantitative structure activityrelationshipmodelsforAmestest129
5.4.2DeeplearningmodelsforAmestest130
5.5Interpretingdeepneuralnetworkmodels134
5.5.1Extractingsubstructures137
5.5.2ComparisonofsubstringswithSARpySAs138
5.5.3ComparisonofsubstructureswithToxtree139
5.6Discussionandconclusions144
5.6.1Afuturefordeeplearningmodels147 References148
6Retrosyntheticspacemodeledbybigdatadescriptors151 ClaudiuN.Lungu
6.1Introduction151
6.2Computer-assistedorganicsynthesis152
6.2.1Retrosyntheticspaceexploredbymoleculardescriptors usingbigdatasets155
6.2.2Theexplorationofchemicalretrosyntheticspaceusing retrosyntheticfeasibilityfunctions156
6.3Quantitativestructure activityrelationshipmodel161
6.4Dimensionalityreductionusingretrosyntheticanalysis164
6.5Discussion166 References167
7Approachinghistoryofchemistrythroughbigdataon chemicalreactionsandcompounds171
GuillermoRestrepo
7.1Introduction171
7.2Computationalhistoryofchemistry172
7.2.1Dataandtools173
7.3Theexpandingchemicalspace,acasestudyforcomputational historyofchemistry178
7.4Conclusions183 Acknowledgments184 References184
8Combinatorialandquantumtechniquesforlargedatasets: hypercubesandhalocarbons187
KrishnanBalasubramanian
8.1Introduction187
8.2Combinatorialtechniquesforisomerenumerationsto generatelargedatasets189
8.2.1Combinatorialtechniquesforlargedatastructures189
8.2.2Mo ¨ biusinversion193
8.2.3Combinatorialresults196
8.3Quantumchemicaltechniquesforlargedatasets198
8.3.1Computationaltechniquesforhalocarbons198
8.3.2Resultsanddiscussionsofquantumcomputationsand toxicityofhalocarbons201
8.4Hypercubesandlargedatasets208
8.5Conclusion211 References212
9Developmentofquantitativestructure activityrelationship modelsbasedonelectrophilicityindex:aconceptualDFT-based descriptor219
RanitaPalandPratimKumarChattaraj
9.1Introduction219
9.2Theoreticalbackground220
9.3Computationaldetails221
9.4Methodology222
9.5Resultsanddiscussion223
9.5.1Tetrahymenapyriformis223
9.5.2Tryphanosomabrucei224
9.6Conclusion226 Acknowledgments226 Conflictofinterest227 References227
10Pharmacophore-basedvirtualscreeningoflargecompound databasescanaid“bigdata”problemsindrugdiscovery231 ApurbaK.Bhattacharjee
10.1Introduction231
10.2Backgroundofdataanalytics,machinelearning,intelligent augmentationmethodsandapplicationsindrugdiscovery233
10.2.1Applicationsofdataanalyticsindrugdiscovery233
10.2.2Machinelearningindrugdiscovery233
10.2.3Applicationofothercomputationalapproachesin drugdiscovery235
10.2.4Predictivedrugdiscoveryusingmolecularmodeling236
10.3Pharmacophoremodeling237
10.3.1Casestudies241
10.4Concludingremarks243 References244
11Anewrobustclassifiertodetecthot-spotsandnull-spotsin protein proteininterface:validationofbindingpocketand identificationofinhibitorsininvitroandinvivomodels247 YanrongJi,XinTong,DanDanXu,JieLiao,RamanaV.Davuluri, Guang-YuYangandRamaK.Mishra
11.1Introduction247
11.2Trainingandtestingoftheclassifier248
11.2.1Variableselectionusingrecursivefeatureelimination249
11.2.2Randomforestperformedbestusingbothpublishedand combineddatasets249
11.3Technicaldetailstodevelopnovelprotein proteininteraction hotspotpredictionprogram251
11.3.1Trainingdata251
11.3.2Buildingandvalidatinganovelclassifierbyevaluating state-of-the-artfeatureselectionandmachinelearning algorithms252
11.4Acasestudy253
11.4.1Identificationofadruggableprotein proteininteraction sitebetweenmutantp53anditsstabilizingchaperone DNAJA1usingourmachinelearning-basedclassifier253
11.4.2BuildingthehomologymodelofDNAJA1and optimizingthemutp53(R175H)structure254
11.4.3Protein proteindocking255
11.4.4Smallmoleculesinhibitorsidentificationthrough drug-likelibraryscreeningagainsttheDNAJA1mutp53R175H interactingpocket256
11.5Discussion259 Authorcontribution260
Acknowledgment260 Conflictsofinterest260 References260
12Miningbigdataindrugdiscovery—triaginganddecisiontrees265
ShahulH.Nilar
12.1Introduction265
12.2Bigdataindrugdiscovery265
12.3Triaging268
12.4Decisiontrees271
12.5Recursivepartitioning271
12.6PhyloGenetic-liketrees273
12.7Multidomainclassification273
12.8Fuzzytreesandclustering276
Acknowledgments278 References278
Section3Bioinformaticsandcomputatioanltoxicology section
13Useofproteomicsdataandproteomics-basedbiodescriptorsinthe estimationofbioactivity/toxicityofchemicalsandnanosubstances285
SubhashC.BasakandMarjanVracko
13.1Introduction285
13.2Proteomicstechnologiesandtheirtoxicologicalapplications286
13.2.1Two-dimensionalgelelectrophoresis286
13.2.2Massspectrometry-basedproteomicstechnologyand theirapplicationsinmathematicalnanotoxicoproteomics290
13.3Discussion292 Acknowledgment295 References295
14Mappinginteractionbetweenbigspaces;activespacefromprotein structureandavailablechemicalspace299 PawanKumar,TaushifKhanandIndiraGhosh
14.1Introduction299
14.2Background301
14.2.1Navigatingproteinfoldspace301
14.2.2Fromaminoacidstringtodynamicstructuralfold301
14.2.3Elementsforclassificationofprotein303
14.2.4Availablemethodsforclassifyingproteins303
14.3Proteintopologyforexploringstructurespace304
14.3.1Modularityinproteinstructurespace305
14.3.2Data-drivenapproachtoextracttopologicalmodule306 x Contents
14.4Scaffoldscurvethefunctionalandcatalyticsites309
14.4.1Signatureofcatalyticsiteinproteinstructures311
14.4.2Proteinfunction-basedselectionoftopologicalspace312 14.4.3Proteindynamicsandtransientsites315 14.4.4Learningmethodsforthepredictionofproteinsand functionalsites316
14.5Proteininteractivesitesanddesigningofinhibitor317 14.5.1Interactionspaceexplorationforenergeticallyfavorable bindingfeaturesidentification317
14.5.2Proteindynamicsguidedbindingfeaturesselection317 14.5.3Proteinflexibilityandexplorationofligandrecognitionsite319 14.5.4Artificialintelligencetounderstandtheinteractionsof proteinandchemical320
14.6Intrinsicallyunstructuredregionsandproteinfunction321 14.7Conclusions322 Acknowledgments323 References323
15Artificialintelligence,bigdataandmachinelearningapproaches ingenome-wideSNP-basedpredictionforprecisionmedicineand drugdiscovery333 IshaJoshi,AnushkaBhrdwaj,RavinaKhandelwal,AditiPande, AnshikaAgarwal,ChillamcherlaDhanalakshmiSrija,RevathyAryaSuresh, ManjuMohan,LimaHazarika,GarimaThakur,TajamulHussain, SarahAlbogami,AnurajNayarisseriandSanjeevKumarSingh 15.1Introduction333
15.2Roleofartificialintelligenceandmachinelearninginmedicine334 15.3Genome-wideSNPprediction339 15.4Artificialintelligence,precisionmedicineanddrugdiscovery340 15.5Applicationsofartificialintelligenceindiseasepredictionand analysisoncology343 15.6Cardiology345
15.7Neurology347 15.8Conclusion348 Abbreviations350 References351
16Applicationsofalignment-freesequencedescriptorsin thecharacterizationofsequencesintheageofbigdata: acasestudywithZikavirus,SARS,MERS,andCOVID-19359 DwaipayanSen,TathagataDey,MarjanVra ˇ cko,AsheshNandyand SubhashC.Basak
16.1Introduction359
16.2Section1—bioinformaticstoday:problemsnow362 16.2.1Whatisbioinformaticsandgenomics?362
16.2.2Annotations362
16.2.3Evolutionofsequencingmethods363
16.2.4Alignment-freesequencedescriptors366
16.2.5Metagenomics367
16.2.6Softwaredevelopment:scenarioandchallenges368
16.2.7Dataformats368
16.2.8Storageandexchange370
16.3Section2—bioinformaticstodayandtomorrow:sustainable solutions370
16.3.1Theneedforbigdata371
16.3.2Softwareanddevelopment373 16.4Summary383 References384
17Scalablequantitativestructure activityrelationshipsystemsfor predictivetoxicology391
SumanK.Chakravarti
17.1Background391
17.2Scalabilityinquantitativestructure activityrelationship modeling393
17.2.1Consequencesofinabilitytoscale394
17.2.2Expandabilityofthetrainingdataset394
17.2.3Efficiencyofdatacuration397
17.2.4Abilitytohandlestereochemistry398
17.2.5Abilitytouseproprietarytrainingdata398
17.2.6Abilitytohandlemissingdata398
17.2.7Abilitytomodifythedescriptorset399
17.2.8Scalingexpertrule-basedsystems399
17.2.9Scalabilityofadverseoutcomepathway-based quantitativestructure activityrelationshipsystems399
17.2.10Scalabilityofthesupportingresources400
17.2.11Scalabilityofquantitativestructure activity relationshipsvalidationprotocols401
17.2.12Scalabilityafterdeployment402
17.2.13Abilitytousecomputerhardwareresourceseffectively402 17.3Summary403 References404
18Frombigdatatocomplexnetwork:anavigationthroughthe mazeofdrug targetinteraction407 ZeWang,MinLi,MuyunTangandGuangHu 18.1Introduction407 18.2Databases409
18.2.1Chemicaldatabases409 18.2.2Databasesfortargets415
18.2.3DatabasesfortraditionalChinesemedicine417
18.3Prediction,construction,andanalysisofdrug targetnetwork418
18.3.1Algorithmstopredictdrug targetinteractionnetwork419
18.3.2Toolsfornetworkconstruction426
18.3.3Networktopologicalanalysis428
18.4Conclusionandperspectives430 Acknowledgments431 References431
19DissectingbigRNA-Seqcancerdatausingmachinelearningto finddisease-associatedgenesandthecausalmechanism437 DipankaTanuSarmah,ShivamKumar,SamratChatterjeeand NandadulalBairagi
19.1Introduction437
19.2Bird’seyeviewoftheanalysisofcancerRNA-Seqdata usingmachinelearning440
19.3Materialsandmethods441
19.3.1Preprocessingofthedata441
19.3.2Featureselection441
19.3.3Classificationlearning442
19.3.4Extractionofdisease-associatedgenes442
19.3.5Validation443
19.4Hand-in-handwalkwithRNA-Seqdata443
19.4.1Datasetselection443
19.4.2Datapreprocessing444
19.4.3Featureselection445
19.4.4Classificationmodel446
19.4.5Identificationofthegenesinvolvedindisease progression447
19.4.6Significanceoftheidentifieddeeplyassociatedgenes447 19.5Conclusion451 References451 Index455
Chemoinformaticsand bioinformaticsbydiscrete mathematicsandnumbers:an adventurefromsmalldatatothe realmofemergingbigdata
SubhashC.Basak DepartmentofChemistryandBiochemistry,UniversityofMinnesotaDuluth,Duluth,MN, UnitedStates
1.1Introduction
“Oh,thethirsttoknow howmany! Thehunger toknow howmany starsinthesky! Wespent ourchildhoodcounting stonesandplants,fingersand toes,grainsofsand,andteeth, ouryouthwaspastcounting petalsandcomets’tails. Wecounted colors,years, lives,andkisses; inthecountry,
oxen;bythesea, thewaves.Ships becameproliferatingciphers.
Numbersmultiplied.”
PabloNeruda,In:Odetonumbers
Acurrentlyemergingtrendinmanyscientificdisciplinesistheirtendencyof beinggraduallytransformed/evolvedintosomeformofinformationscience(Basak etal.,2015;DehmerandBasak,2012;Kerberetal.,2014).Intherealmofchemoinformaticsandbioinformatics,inparticular,methodsofdiscretemathematics likegraphtheory,networktheory,informationtheoryetc.aregainingmomentum asusefultoolsintherepresentation,characterization,andcomparisonofmolecular andbiologicalsystemsandtheirstructuresaswellasinthepredictionofproperty/ bioactivity/toxicityofchemicalsfornewdrugdiscoveryandenvironmentalprotection(Basak,1987,2010,2013a,2014;Basaketal.,1988b,2015;Baydaetal., 2019;Bielinska-Wazetal.,2007;Bragaetal.,2018;Chakravarti,2021;Ciallella andZhu,2019;Diudeaetal.,2018;Ginietal.,2013;Guoetal.,2001;Kerber etal.,2014;Khanetal.,2018;Nandy,2015;KierandHall,1986,1999;Nandy etal.,2006;Osolodkinetal.,2015;Randicetal.,2000,2001,2004,2011;Restrepo andVillaveces,2013;Rouvray,1991;Sabirovetal.,2021;ToropovandToropova, 2021;Vra ˇ ckoetal.,2018,2021a,b;Wangetal.,2021;Winkleretal.,2014).
Theimpetusforthedevelopmentofchemoinformaticsandbioinformaticstools/ methodshascomefromdifferentdirections.Innewdrugdesign,thousandsofderivativesoftheinitiallydiscovered“lead”compoundhavetobesynthesizedand testedinordertofindoneusefuldrug.Thisjourneyoftheleadfromthechemist’s desktothebedsideofthepatientinvolvesaspanofabout10yearsandanexpenditureofoverUS$2billion(DiMasietal.,2016).Synthesisandtestingofallpossible chemicalderivativesoftheidentifiedleadcompoundisprohibitivelycostly.Under suchcircumstancesinsilicoapproachesofchemoinformaticscangiveusfastand cost-effectiveestimationofpropertiesofpromisingderivativesoftheleadchemicalsnecessaryforthepredictionofthemostprobablepharmacologicalandtoxicologicalprofiles(Table1.1).Thus,chemoinformaticstoolscanassistthedrug designerasa decisionsupportsystem.Ithasbeennotedthatcurrentlynodrugis developedwithoutthepriorevaluationbyquantitativestructure activityrelationship(QSAR)methods(Santos-Filhoetal.,2009).
TheToxicSubstancesControlAct(TSCA,2021)Inventory,maintainedbythe UnitedStatesEnvironmentalProtectionAgency(USEPA),currentlyhasmorethan 86,000chemicals.MostoftheTSCAchemicalshaveverylittleornoexperimental datarequiredfortheirtoxicityestimation.Detailedlaboratorytestingofallthese chemicalsandtheirpossiblemetabolitesproducedintheexposedorganismsincludinghumanswouldbeprohibitivelycostly.Inthefaceofthislackofavailabledata, twoapproachesareusedbytheregulatoryagencies:(a)class-specificQSARmodelsand(b)quantitativemolecularsimilarityanalysis(QMSA)-basedmodelingof 4BigDataAnalyticsinChemoinformaticsandBioinformatics
Table1.1 Apartiallistofimportantphysical,pharmacological,andtoxicological propertiesprerequisitetotheevaluationofchemicalsfornewdrugdiscoveryand environmentalprotection.
PhysicochemicalPharmacological/toxicological
Molarvolume
Macromoleculelevel
BoilingpointReceptorbinding(KD)
MeltingpointMichaelisconstant(Km)
VaporpressureInhibitorconstant(Ki)
WatersolubilityDNAalkylation
Dissociationconstant(pKa)UnscheduledDNAsynthesis
Partitioncoefficient
Celllevel
Octanol-water(logP)Salmonellamutagenicity
Air-waterMammaliancelltransformation
Sediment-water
Organismlevel(acute)
Reactivity(electrophile)Algae
Invertebrates
Fish Birds
Mammals
Organismlevel(chronic)
Bioconcentration
Carcinogenicity
Reproductivetoxicity
Delayedneurotoxicity
Biodegradation
propertiesusingstructuralanalogs(Aueretal.,1990).Thesituationbecomesmore numerousandcomplexifoneconsidersthebiotransformationandpharmacokinetic dataofthechemicals(TSCAmetabolismandpharmacokinetics,2021).Asimilar situationexistsintheEuropeanUnionwiththechemicalsincommerce(European ChemicalsAgency,2021)listshowingmorethan100,000chemicalsregistered withthesystem.
Table1.1 providesapartiallistofphysicochemical,pharmacological,andtoxicologicalpropertiesthatdrugdesignersandriskassessorsofchemicalsfrequently useinevaluatingtheirbeneficialanddeleteriouseffects(Basaketal.,1990).
Engenderedallthatbeinghath.
Andthoughtheyseemtoclingtogether, Andform“associations”here, Yet,soonorlate,theybursttheirtether, Andthroughthedepthsofspacecareer.”
—JamesClerkMaxwell
ThecurrentQSARparadigmdidnotariseoutofoneorafew“aha”moments, butitemergedthroughtheconfluenceofadiversesetofideasoriginatedbyquitea fewresearchersofdifferentdisciplinesoverthepastcoupleofcenturies.Fora recentreview,pleasesee Basak(2021a).Someseminalaspectsofthedevelopments ofmodernchemoinformaticsarediscussedasfollows.
1.2.1Physicalproperty:colligativeversusconstitutive
“Inordertodescribeanaspectofholisticrealitywehavetoignorecertainfactors suchthattheremainderseparatesintofacts.Inevitably,suchadescriptionistrue onlywithintheadoptedpartitionoftheworld,thatis,withinthechosencontext.”
—Hans Primas(1981),Chemistry,QuantumMechanicsandReductionism
Inphysicalchemistry,a colligativeproperty,forexample,loweringofvapor pressure,elevationofboilingpoint,depressionoffreezingpoint,andosmoticpressure,ofsolutionsisapropertythatdependssolelyupontheconcentrationofsolute moleculesorions,beingindependentoftheconstitutionoridentityofthesolute. Constitutiveproperty,ontheotherhand,dependsontheconstitutionorstructureof thesubstance.The AmericanHeritageDictionary oftheEnglishLanguage,5th Edition,statesthefollowingregardingthewordconstitutive:
“Inphysicalchemistry,atermintroducedbyOstwaldtodenotethosepropertiesof acompoundwhichdependontheconstitutionofthemolecule,oronthemodeof unionandarrangementoftheatomsinthemolecule.”
1.2.2Earlybiochemicalobservationsontherelationship betweenchemicalstructureandbioactivityofmolecules
Foralmostacentury,variousresearchersinbiochemistryandpharmacologygenerateddataontherelationbetweenthestructureofmoleculesandtheirbioactivities. Mostprobablyoneoftheearliestwasthe1928findingof QuastelandWooldridge (1928) thatmalonicacidcompetitivelyinhibitedtheactivityoftheKrebscycle enzymesuccinicdehydrogenase.Althoughthesubstratesuccinicacidandthe 6BigDataAnalyticsinChemoinformaticsandBioinformatics
inhibitormalonicaciddifferedbyonemethylene( CH2)group,thecatalyticsite oftheenzymestillrecognizedmalonicacid.Thisseminalobservationmaybe lookeduponastherationalbasisforthesynthesisofanalogsofnucleicacidbases forcancerchemotherapy(HitchingsandElion,1954)andthemoremodernchemoinformaticsapproachtocomputer-aideddrugdesignusingtheconceptofpharmacophore(Bhattacharjee,2015).Theantibioticpenicillininhibitscellwall biosynthesisinbacteriabyinterferingwiththetranspeptidationreactionresponsible forthecrosslinkingofmucopeptidechainsinthecellwallpolymer.ThisisattributedtoitsputativestructuralsimilaritytotheD-alanyl-D-alanineportionofthe peptidechain(GoodmanandGilman,1990).
1.2.3Linearfreeenergyrelationship:themultiparameterHansch approachtoquantitativestructure activityrelationship
Asdescribedby HanschandLeo(1979),intheearly1900stheEnglishschoolof organicchemists(Ingold,1953)becameinterestedinthemechanismsofreactions oforganicmolecules.Oneapproachwastomakeasetofstructuralmodifications inaparentmoleculeandthenobservetheeffectsofthesubstitutionsontheratesor equilibriaofareactionwithareactantunderstandardconditions.Onecoulddraw conclusionsabouttheelectronicandstericrequirementsofagivenreactionfrom theanalysisoftheperturbationsofthereactioncenterbythesubstituents.Theproblemsofapplicationsoftheseconcepts,asindicatedby HanschandLeo(1979), were:
“Thedifficultywiththeseearlyandimportantideaswasthatnonumericalscales wereavailablethatcouldbeusedtoquantifyeachoftheseeffectsthatcould operatesinglyorinconcert. Evenwhensuchscaleshadbeendevised,itwas difficulttomakeprogressintheseparationofsubstituenteffectsbeforehighspeedcomputersbecamegenerallyavailable (approximately1960).”
Oneimportantbreakthroughinthefieldofmechanisticorganicchemistrycame when Hammett(1937) proposedthenowwell-knownHammettequation.He definedtheparameter σ asfollows:
5 log
(1.1) where KH istheionizationconstantforbenzoicacidinwaterat25 Cand Kx isthe ionizationconstantforitsmetaorparaderivativeunderthesameexperimentalconditions.Positivevaluesof σ indicateelectronwithdrawalbythesubstituentfrom thearomaticringandnegativevaluesrepresentelectronreleasefromthesubstituent tothering.
Inthesecondhalfofthe20thcentury, Taft(1952) formulatedthelinearfree energy-relatedstericdescriptor Es
Themultiparameterlinearfreeenergyrelationship(LFER)approach,popularly knownasthe“HanschAnalysis,”toquantitativestructure property activity
relationship(QSPR/QSAR),derivedfromphysicalorganicchemistry,attemptedto predictproperty/bioactivityofmoleculesusingacombinationoftheirelectronic, steric,andhydrophobicparameters(HanschandLeo,1995):
In Eq.(1.2),BAstandsforbiologicalactivity,log P standsforthelogarithmof thepartitioncoefficient(experimentallydeterminedorcalculatedfromstructure)of thechemical, σ usuallyrepresents Hammett’s(1937) electronicdescriptor,and Es usuallysymbolizesTaft’sstericparameter(Taft,1952).AperusalofLEFR-based QSARmodelswouldindicatethatdifferentvarietiesofhydrophobic,steric,and electronicparametershavebeendevelopedandusedinnumerouscorrelationstudies(HanschandLeo,1995).Ashortdescriptionofthehistoricaltimelineforthe evolutionoftheLFERapproachisdepictedin Fig.1.1.
TheLFERapproachgivesgoodpredictivemodelsforcongenericsetsofmolecules.Asdiscussedabove,bothfordrugdesignandhazardassessmentofchemicals
Figure1.1 Ashorthistory(1868-date)ofthedevelopmentoflinearfreeenergyrelationship approachforquantitativestructure activityrelationshipmodelingbasedonphysical propertiesandsubstituentconstantsderivedfromphysicalorganicchemistry.Formore informationpleasesee: Basak(2013a,2021a) and HanschandLeo(1995).Inthisapproach, aproperty(P1)ofamoleculeisestimatedfromanotheravailableproperty(P2)ora combinationofotherproperties.
Hansch approach 1962
bioactivity = f (Steric, electronic & hydrophobic parameters)
Hammett sigma 1937
Crum-Brown & Fraser 1868: Prop= f (size, complexity)
Taft steric parameter 1952
LFER Prop-prop correlation approach: P1 =f(P2)
Overton (1896) ; Meyer (1899)
Narcosis = f (oil-water partitioncoefficient)
Chemoinformaticsandbioinformaticsbydiscretemathematicsandnumbers
weneedtoestimatethepropertiesandbioactivitiesofchemicalswhicharestructurallydiverse(BasakandMajumdar,2016).Sometimes,onecouldwishtoestimate pharmacologicalandtoxicologicalprofilesofchemicalsnotyetsynthesized. ModelsbasedontheLFER-typeexperimentaldataareoflittleutilityinsuchcases. Furthermore,forapplicationsinchemicalengineeringandtechnologicalprocesses weneedtoknowthevaluesofmanypropertiesofsubstances(Drefahland Reinhard,1998;Lymanetal.,1990).Theuseofgoodqualityexperimentalproperty valuesarealwaysdesirable,butsuchdataareoftenunavailable.TheuseofQSARpredictedpropertiesutilizingcomputeddescriptorsastheindependentvariablesis generallythepracticalalternative(Katritzkyetal.,1995,2001).Morerecently, manylargeanddiversedatabasesofpropertiesneededfordrugdesignandpredictivetoxicologyarebecomingavailableinthepublicdomain.Theseareresources availableforthedevelopmentofbroad-basedmodelsforproperty/bioactivityestimation(Gadaletaetal.,2019;Mansourietal.,2018;Mengetal.,2021).
Duringthesecondhalfofthe20thcenturyandthefirstquarterofthiscentury, variouschemoinformaticsapproacheshavegivenusmoleculardescriptorswhich canbecomputeddirectlyfromthemolecularstructurewithouttheinputofany otherexperimentaldata.Suchdescriptorsarefindingsidespreadapplicationsinthe formulationofusefulQSARmodels(Basak,2021a,2012b,2013a,2014;Drefahl andReinhard,1998;Katritzkyetal.,1995,2001;KierandHall,1986,1999).
1.2.4Chemicalgraphtheoryandquantumchemistryasthe sourceofchemodescriptors
“Byconventionsweetandbyconventionbitter,byconventionhot,byconvention cold,byconventioncolor;butinrealityatomsandvoid.”
—Democritus
“Thefundamentallawsnecessaryforthemathematicaltreatmentofalargepartof physicsandthewholeofchemistryarethuscompletelyknown,andthedifficulty liesonlyinthefactthatapplicationoftheselawsleadstoequationsthataretoo complextobesolved.”
—PaulDirac
1.2.4.1Topologicalindices—graphtheoreticdefinitionsand calculationmethods
Agraph, G,isdefinedasanorderedpairconsistingoftwosets V and R, G 5 [V (G), R],where V(G)representsafinitenonemptysetofpoints,and R isabinary relationdefinedontheset V(G).Theelementsof V arecalledverticesandtheelementsof R,alsosymbolizedby E(G)or E,arecallededges.Suchanabstractgraph iscommonlyvisualizedbyrepresentingelementsof V(G)aspointsandbyconnectingeachpair(u, v)ofelementsof V(G)withalineifandonlyif(u, v)ER.Thevertex, v,andedge, e,areincidentwitheachother,asare u and e.Twovertices u and
v in G arecalledadjacentif(u, v)ER,thatis,theyareconnectedbyanedge.Awalk ofagraphisasequencebeginningandendingwithverticesinwhichverticesand edgesalternateandeachedgeisincidentwithverticesimmediatelyprecedingand followingit.Awalkoftheform v0, e1, v1, e2, ..., vn joinsvertices v0 and vn.The lengthofawalkisthenumberofedgesinthewalk.Awalkisclosedif v0 5 vn, otherwiseitisopen.Aclosedwalkwithnpointsisacycleifallitspointsaredistinctand n $ 3.Apathisanopenwalkinwhichallverticesaredistinct.Agraph G isconnectedifeverypairofitsverticesisconnectedbyapath.Agraph G isamultigraphifitcontainsmorethanoneedgebetweenatleastonepairofadjacentvertices,otherwise, G isasimplegraph.Thedistance d (u, v)betweenvertices u and v in G isthelengthoftheshortestpathconnecting u and v.
Becauseofthegeneralnatureofgraph-theoretic(GT)methodsintherepresentationofobjectsthismethodhasbeenusedinsuchdiverseareasastheoreticalphysics,chemistry,biologicalandsocialsciences,engineering,computerscienceand linguistics(Harary,1986).Forexample,GThasbeenusedintherepresentationand comparisonofproteins,characterizationofthenucleotidesequencetopologyin DNAandRNAsequences(Nandy,2015;Nandyetal.,2006;Randicetal.,2000, 2011),representationofproteinspotsofproteomicsmaps(Randi ´ cetal.,2001), foldingpatternsinproteinstructures(Khanetal.,2018;Liuetal.,2006),structural characterizationofnanosubstances(ToropovandToropova,2021),tonamejusta few.
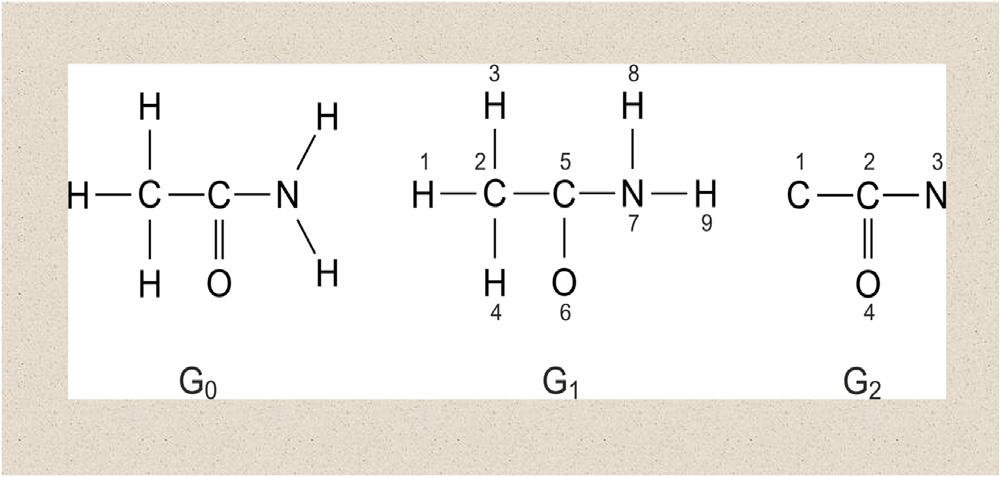
Forchemicalgraphtheoryresearchandapplications(Basak,2013a;Basaketal., 2011;Janezicetal.,2015),amoleculargraphrepresentsmoleculartopologywhere V representsthesetofatomsand E usuallysymbolizesthesetofcovalentbonds presentinthemolecule.Itshouldbenoted,however,thattheset E shouldnotbe limitedtocovalentbondsonly.Infact,elementsof E maysymbolizeanytypeof bond,viz.,covalent,ionic,orhydrogenbonds,etc.Itwasemphasizedby Basak etal.(1988a) thatweightedpseudographsconstituteaveryversatilemodelforthe representationofawiderangeofchemicalspecies. Fig.1.2 depictsthechemical structure,labeledhydrogen-filledgraphandlabeledhydrogen-suppressedgraphof themoleculeacetamide.Itmaybementionedherethatalargenumberofmolecules
Figure1.2 Structuralformula(G0),labeledhydrogen-filledgraph(G1),andlabeled hydrogen-suppressedgraph(G2)ofacetamide.